 
Available fields are: Gene name ( HGNC name or synonyms), Protein class, Chromosome, External identifier ( Ensembl gene, transcript or protein identifier, UniProt accession number, NCBI Entrez gene identifier), Subcellular location based on immunofluorescent staining in three different cell lines, Organ-, Tissue-, Cancer- and Cell line expression, Antibody validation results in four different assays ( IH=immunohistochemistry, IF=immunoflourescence, WB=Western blot, PA=Protein array), evidence scores and a filtration on Genes with antibodies only and Genes with knowledge-based annotated protein expression ( IH, IF). A fourth modification of the protein backbone is the hydrogenation of dehydroamino acids to afford D-amino acids catalyzed by NAD(P)H-dependent reductases. When the amino group of an amino acid combines with the carboxyl group of another amino. Several citrullinated proteins are associated with RA, suggesting that ACPAs may react with different sequence patterns, separating them from traditional. Specific fields can be searched by using the "Fields"-function. Why Peptide bonds are Backbone of the Proteins Formation of a Peptide bonds. The beta sheet, (-sheet) (also -pleated sheet) is a common motif of the regular protein secondary structure.Beta sheets consist of beta strands (-strands) connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. 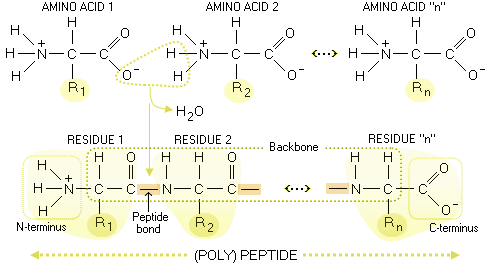
The free text query word needs to be at least three characters long. The free text search will scan for complete and partial matches to gene names, gene synonyms, gene descriptions, external ( UniProt, Ensembl, NCBI Entrez Gene) gene and protein identifiers, protein classes, Gene Ontology identifiers and descriptions, antibody identifiers and image annotations. A protein ultimately folds in space to form a unique 3D shape, which usually contains some alpha helices and beta sheets. The polypeptidc chain of a natural protein may be called a normal-all-L chain normal defining the direc- tion of peptide bonds along the chain, and all-t. 
We must consider the effects of the 20 different R groups (side chains) which complicate the folding process. Secondary structure elements typically spontaneously form as an intermediate before the. Protein folding is determined by much more than the formation of hydrogen bonds between backbone donors and acceptors. Cyclization is commonly employed in efforts to improve the target binding affinity of peptide-based probes and therapeutics. 1 The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well. The search function can be used for free text search (type anything in the search field), or for more complex queries using "Fields" (see examples). Protein secondary structure is the local spatial conformation of the polypeptide backbone excluding the side chains. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
